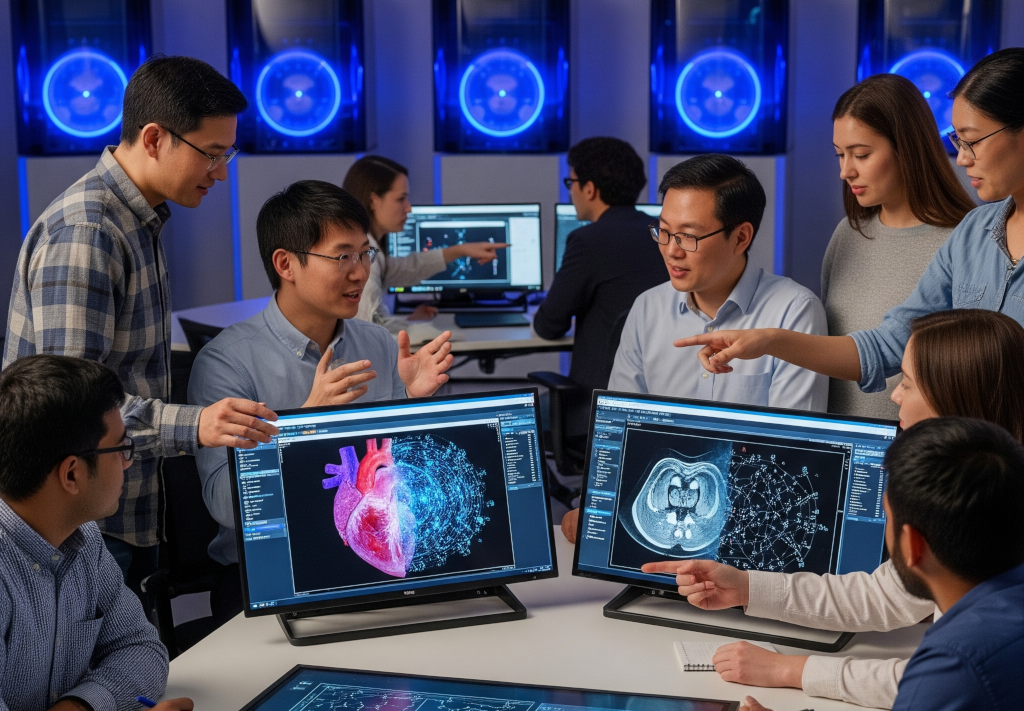
Computational Healthcare & Oncology Informatics (CHOI) Lab at Sidney Kimmel Medical College, Thomas Jefferson University, led by Wookjin Choi, Ph.D.
We develop interpretable AI for medical imaging — quantitative image analysis, radiomics, and deep learning for cancer diagnosis, radiation therapy planning, and outcome prediction.
Focus areas: Lung cancer screening · Cardiotoxicity prediction (Cardiac PET radiomics) · MR-guided adaptive radiotherapy (MRI-Linac) · Pancreatic / hepatic / head-and-neck cancer imaging · Interpretable deep learning (PathCNN, CIRDataset, Spiculation).